The Basics#
PyAutoFit is a Python based probabilistic programming language for model fitting and Bayesian inference of large datasets.
The basic PyAutoFit API allows us a user to quickly compose a probabilistic model and fit it to data via a log likelihood function, using a range of non-linear search algorithms (e.g. MCMC, nested sampling).
This overview gives a run through of:
Models: Use Python classes to compose the model which is fitted to data.
Instances: Create instances of the model via its Python class.
Analysis: Define an
Analysisclass which includes the log likelihood function that fits the model to the data.Searches: Choose an MCMC, nested sampling or maximum likelihood estimator non-linear search algorithm that fits the model to the data.
Model Fit: Fit the model to the data using the chosen non-linear search, with on-the-fly results and visualization.
Results: Use the results of the search to interpret and visualize the model fit.
This overviews provides a high level of the basic API, with more advanced functionality described in the following overviews and the PyAutoFit cookbooks.
Example#
To illustrate PyAutoFit we’ll use the example modeling problem of fitting a 1D Gaussian profile to noisy data.
To begin, lets import autofit (and numpy) using the convention below:
import autofit as af
import numpy as np
The example data with errors (black) is shown below:
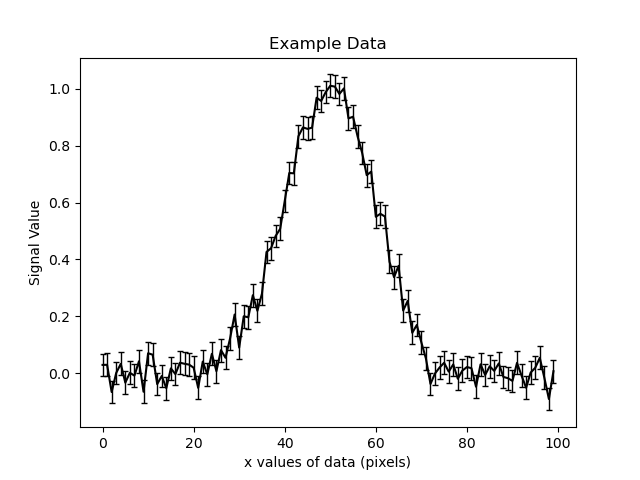
The 1D signal was generated using a 1D Gaussian profile of the form:
Where:
x: The x-axis coordinate where theGaussianis evaluated.
N: The overall normalization of the Gaussian.
sigma: Describes the size of the Gaussian.
Our modeling task is to fit the data with a 1D Gaussian and recover its parameters (x, N, sigma).
Model#
We therefore need to define a 1D Gaussian as a PyAutoFit model.
We do this by writing it as the following Python class:
class Gaussian:
def __init__(
self,
centre=0.0, # <- PyAutoFit recognises these constructor arguments
normalization=0.1, # <- are the Gaussian`s model parameters.
sigma=0.01,
):
"""
Represents a 1D `Gaussian` profile, which can be treated as a
PyAutoFit model-component whose free parameters (centre,
normalization and sigma) are fitted for by a non-linear search.
Parameters
----------
centre
The x coordinate of the profile centre.
normalization
Overall normalization of the `Gaussian` profile.
sigma
The sigma value controlling the size of the Gaussian.
"""
self.centre = centre
self.normalization = normalization
self.sigma = sigma
def model_data_1d_via_xvalues_from(self, xvalues: np.ndarray) -> np.ndarray:
"""
Returns the 1D Gaussian profile on a line of Cartesian x coordinates.
The input xvalues are translated to a coordinate system centred on the
Gaussian, by subtracting its centre.
The output is referred to as the `model_data` to signify that it is
a representation of the data from the model.
Parameters
----------
xvalues
The x coordinates for which the Gaussian is evaluated.
"""
transformed_xvalues = xvalues - self.centre
return np.multiply(
np.divide(self.normalization, self.sigma * np.sqrt(2.0 * np.pi)),
np.exp(-0.5 * np.square(np.divide(transformed_xvalues, self.sigma))),
)
The PyAutoFit model above uses the following format:
The name of the class is the name of the model, in this case, “Gaussian”.
The input arguments of the constructor (the
__init__method) are the parameters of the model, in this casecentre,normalizationandsigma.The default values of the input arguments define whether a parameter is a single-valued
floator a multi-valuedtuple. In this case, all 3 input parameters are floats.It includes functions associated with that model component, which are used when fitting the model to data.
To compose a model using the Gaussian class above we use the af.Model object.
model = af.Model(Gaussian)
print("Model ``Gaussian`` object: \n")
print(model)
This gives the following output:
Model `Gaussian` object:
Gaussian (centre, UniformPrior [1], lower_limit = 0.0, upper_limit = 100.0),
(normalization, LogUniformPrior [2], lower_limit = 1e-06, upper_limit = 1000000.0),
(sigma, UniformPrior [3], lower_limit = 0.0, upper_limit = 25.0)
Note
You may be wondering where the priors above come from. PyAutoFit allows a user to set up configuration files that define the default priors on every model parameter, to ensure that priors are defined in a consistent and concise way. More information on configuration files is provided in the next overview and the cookbooks.
The model has a total of 3 parameters:
print(model.total_free_parameters)
All model information is given by printing its info attribute:
print(model.info)
This gives the following output:
Total Free Parameters = 3
model Gaussian (N=3)
centre UniformPrior [1], lower_limit = 0.0, upper_limit = 100.0
normalization LogUniformPrior [2], lower_limit = 1e-06, upper_limit = 1000000.0
sigma UniformPrior [3], lower_limit = 0.0, upper_limit = 25.0
The priors can be manually altered as follows:
model.centre = af.UniformPrior(lower_limit=0.0, upper_limit=100.0)
model.normalization = af.UniformPrior(lower_limit=0.0, upper_limit=1e2)
model.sigma = af.UniformPrior(lower_limit=0.0, upper_limit=30.0)
Printing the model.info displayed these updated priors.
print(model.info)
This gives the following output:
Total Free Parameters = 3
model Gaussian (N=3)
centre UniformPrior [4], lower_limit = 0.0, upper_limit = 100.0
normalization UniformPrior [5], lower_limit = 0.0, upper_limit = 100.0
sigma UniformPrior [6], lower_limit = 0.0, upper_limit = 30.0
Instances#
Instances of a PyAutoFit model (created via af.Model) can be created, where an input vector of parameter
values is mapped to create an instance of the model’s Python class.
We first need to know the order of parameters in the model, so we know how to define the input vector. This
information is contained in the models paths attribute:
print(model.paths)
This gives the following output:
[('centre',), ('normalization',), ('sigma',)]
We input values for the 3 free parameters of our model following the order of paths above (centre=30.0, normalization=2.0 and sigma=3.0):
instance = model.instance_from_vector(vector=[30.0, 2.0, 3.0])
This is an instance of the Gaussian class.
print("Model Instance: \n")
print(instance)
This gives the following output:
Model Instance:
<__main__.Gaussian object at 0x7f3e37cb1990>
It has the parameters of the Gaussian with the values input above.
print("Instance Parameters \n")
print("x = ", instance.centre)
print("normalization = ", instance.normalization)
print("sigma = ", instance.sigma)
This gives the following output:
Instance Parameters
x = 30.0
normalization = 2.0
sigma = 3.0
We can use functions associated with the class, specifically the model_data_1d_via_xvalues_from function, to
create a realization of the Gaussian and plot it.
xvalues = np.arange(0.0, 100.0, 1.0)
model_data = instance.model_data_1d_via_xvalues_from(xvalues=xvalues)
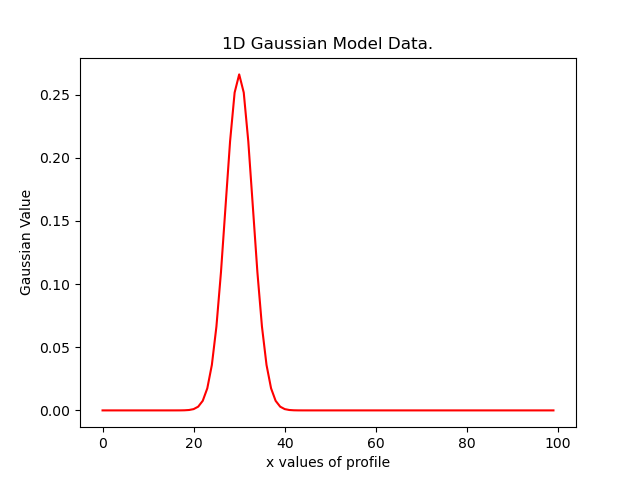
plt.plot(xvalues, model_data, color="r")
plt.title("1D Gaussian Model Data.")
plt.xlabel("x values of profile")
plt.ylabel("Gaussian Value")
plt.show()
plt.clf()
Here is what the plot looks like:
Note
Mapping models to instance of their Python classes is an integral part of the core PyAutoFit API. It enables the advanced model composition and results management tools illustrated in the following overviews and cookbooks.
Analysis#
We now tell PyAutoFit how to fit the model to data.
We define an Analysis class, which includes:
An
__init__constructor, which takes as input thedataandnoise_map(this can be extended with anything else necessary to fit the model to the data).A
log_likelihood_function, defining how for aninstanceof the model we fit it to the data and return a log likelihood value.
Read the comments and docstrings of the Analysis object below in detail for a full description of how an analysis works.
class Analysis(af.Analysis):
def __init__(self, data: np.ndarray, noise_map: np.ndarray):
"""
The ``Analysis`` class acts as an interface between the data and
model in **PyAutoFit**.
Its ``log_likelihood_function`` defines how the model is fitted to
the data and it is called many times by the non-linear search fitting
algorithm.
In this example the ``Analysis`` ``__init__`` constructor only contains
the ``data`` and ``noise-map``, but it can be easily extended to
include other quantities.
Parameters
----------
data
A 1D numpy array containing the data (e.g. a noisy 1D signal)
fitted in the readthedocs and workspace examples.
noise_map
A 1D numpy array containing the noise values of the data, used
for computing the goodness of fit metric, the log likelihood.
"""
super().__init__()
self.data = data
self.noise_map = noise_map
def log_likelihood_function(self, instance) -> float:
"""
Returns the log likelihood of a fit of a 1D Gaussian to the dataset.
The data is fitted using an ``instance`` of the ``Gaussian`` class
where its ``model_data_1d_via_xvalues_from`` is called in order to
create a model data representation of the Gaussian that is fitted to the data.
"""
"""
The ``instance`` that comes into this method is an instance of the ``Gaussian``
model above, which was created via ``af.Model()``.
The parameter values are chosen by the non-linear search and therefore are based
on where it has mapped out the high likelihood regions of parameter space are.
The lines of Python code are commented out below to prevent excessive print
statements when we run the non-linear search, but feel free to uncomment
them and run the search to see the parameters of every instance
that it fits.
# print("Gaussian Instance:")
# print("Centre = ", instance.centre)
# print("Normalization = ", instance.normalization)
# print("Sigma = ", instance.sigma)
"""
"""
Get the range of x-values the data is defined on, to evaluate the model of the Gaussian.
"""
xvalues = np.arange(self.data.shape[0])
"""
Use these xvalues to create model data of our Gaussian.
"""
model_data = instance.model_data_1d_via_xvalues_from(xvalues=xvalues)
"""
Fit the model gaussian line data to the observed data, computing the residuals,
chi-squared and log likelihood.
"""
residual_map = self.data - model_data
chi_squared_map = (residual_map / self.noise_map) ** 2.0
chi_squared = sum(chi_squared_map)
noise_normalization = np.sum(np.log(2 * np.pi * self.noise_map**2.0))
log_likelihood = -0.5 * (chi_squared + noise_normalization)
return log_likelihood
Create an instance of the Analysis class by passing the data and noise_map.
analysis = Analysis(data=data, noise_map=noise_map)
Non Linear Search#
We now have a model to fit to the data, and an analysis class that performs this fit.
We now choose our fitting algorithm, called the “non-linear search”, and fit the model to the data.
PyAutoFit supports many non-linear searches, which broadly speaking fall into three categories, MCMC, nested sampling and maximum likelihood estimators.
For this example, we choose the nested sampling algorithm Dynesty.
search = af.DynestyStatic(
nlive=100,
)
Note
The default settings of the non-linear search are specified in the configuration files of PyAutoFit, just like the default priors of the model components above. More information on configuration files is provided in the next overview and the cookbooks.
Model Fit#
We begin the non-linear search by passing the model and analysis to its fit method.
print(
The non-linear search has begun running.
This Jupyter notebook cell with progress once the search
has completed - this could take a few minutes!
)
result = search.fit(model=model, analysis=analysis)
print("The search has finished run - you may now continue the notebook.")
Result#
The result object returned by the fit provides information on the results of the non-linear search.
The info attribute shows the result in a readable format.
print(result.info)
The output is as follows:
Bayesian Evidence 167.54413502
Maximum Log Likelihood 183.29775793
Maximum Log Posterior 183.29775793
model Gaussian (N=3)
Maximum Log Likelihood Model:
centre 49.880
normalization 24.802
sigma 9.849
Summary (3.0 sigma limits):
centre 49.88 (49.51, 50.29)
normalization 24.80 (23.98, 25.67)
sigma 9.84 (9.47, 10.25)
Summary (1.0 sigma limits):
centre 49.88 (49.75, 50.01)
normalization 24.80 (24.54, 25.11)
sigma 9.84 (9.73, 9.97)
Results are returned as instances of the model, as illustrated above in the instance section.
For example, we can print the result’s maximum likelihood instance.
print(result.max_log_likelihood_instance)
print("\nModel-fit Max Log-likelihood Parameter Estimates: \n")
print("Centre = ", result.max_log_likelihood_instance.centre)
print("Normalization = ", result.max_log_likelihood_instance.normalization)
print("Sigma = ", result.max_log_likelihood_instance.sigma)
This gives the following output:
Model-fit Max Log-likelihood Parameter Estimates:
Centre = 49.87954357347897
Normalization = 24.80227227310798
Sigma = 9.84888033338011
A benefit of the result being an instance is that we can use any of its methods to inspect the results.
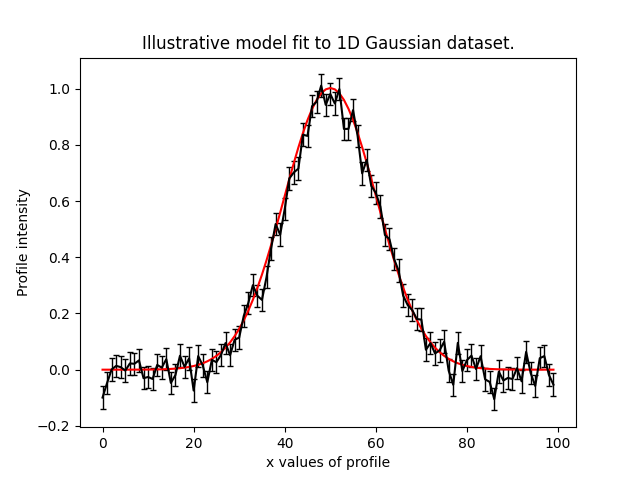
Below, we use the maximum likelihood instance to compare the maximum likelihood Gaussian to the data.
model_data = result.max_log_likelihood_instance.model_data_1d_via_xvalues_from(
xvalues=np.arange(data.shape[0])
)
plt.errorbar(
x=xvalues, y=data, yerr=noise_map, color="k", ecolor="k", elinewidth=1, capsize=2
)
plt.plot(xvalues, model_data, color="r")
plt.title("Dynesty model fit to 1D Gaussian dataset.")
plt.xlabel("x values of profile")
plt.ylabel("Profile normalization")
plt.show()
plt.close()
The plot appears as follows:
Samples#
The results object also contains a Samples object, which contains all information on the non-linear search.
This includes parameter samples, log likelihood values, posterior information and results internal to the specific algorithm (e.g. the internal dynesty samples).
This is described fully in the results overview, below we use the samples to plot the probability density function corner of the results.
plotter = aplt.NestPlotter(samples=result.samples)
plotter.corner()
The plot appears as follows:

Wrap Up#
This overview explains the basic PyAutoFit. It used a simple model, a simple dataset, a simple model-fitting problem and pretty much the simplest parts of the PyAutoFit API.
You should now have a good idea for how you would define and compose your own model, fit it to data with a chosen non-linear search, and use the results to interpret the fit.
The PyAutoFit API introduce here is very extensible and very customizable, and therefore easily adapted to your own model-fitting problems.
The next overview describes how to use PyAutoFit to set up a scientific workflow, in a nutshell how to use the API to make your model-fitting efficient, manageable and scalable to large datasets.
Resources#
The autofit_workspace: has numerous examples of how to perform more complex model-fitting tasks:
The following cookbooks describe how to use the API for more complex model-fitting tasks:
Model Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/model.html
Compose complex models from multiple Python classes, lists, dictionaries and customize their parameterization.
Analysis Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/search.html
Customize the analysis with model-specific output and visualization.
Searches Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/analysis.html
Choose from a variety of non-linear searches and customize their behaviour, for example outputting results to hard-disk and parallelizing the search.
Results Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/result.html
The variety of results available from a fit, including parameter estimates, error estimates, model comparison and visualization.
Configs Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/configs.html
Customize default settings using configuration files, for example setting priors, search settings and visualization.
Multiple Dataset Cookbook: https://pyautofit.readthedocs.io/en/latest/cookbooks/multiple_datasets.html
Fit multiple datasets simultaneously, by combining their analysis classes such that their log likelihoods are summed.